Radford University
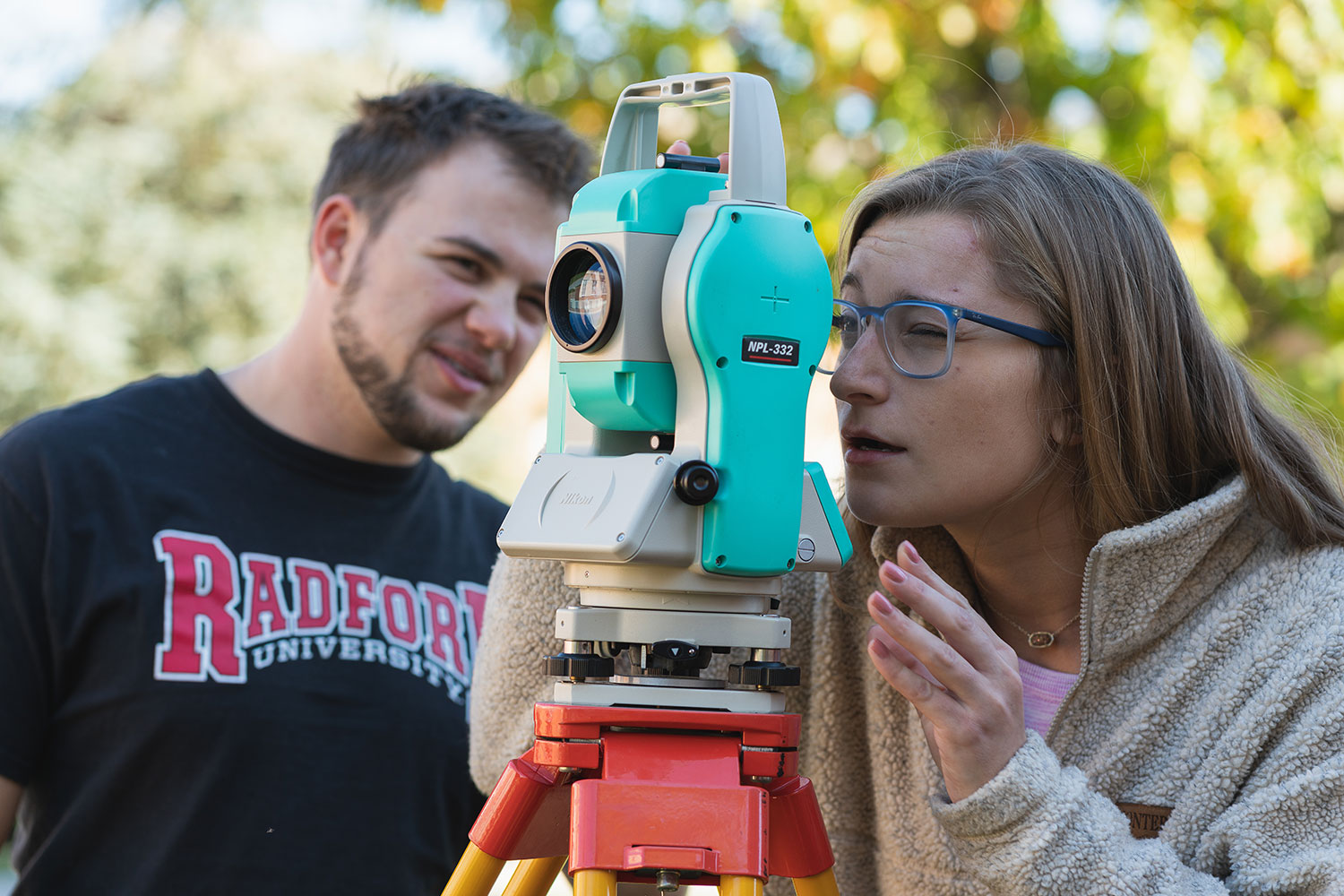
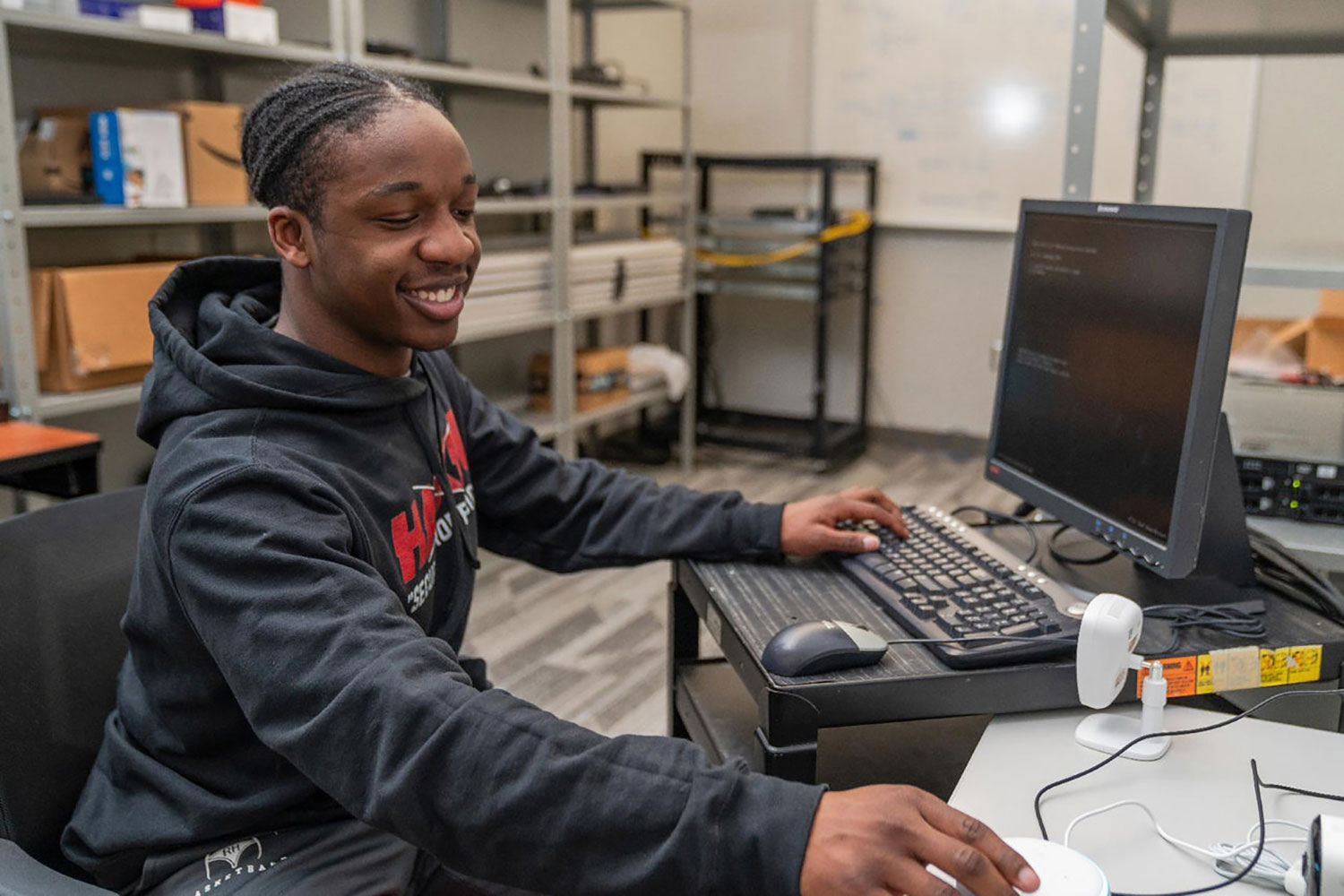
Artis College of Science and Technology
Innovation and Discovery
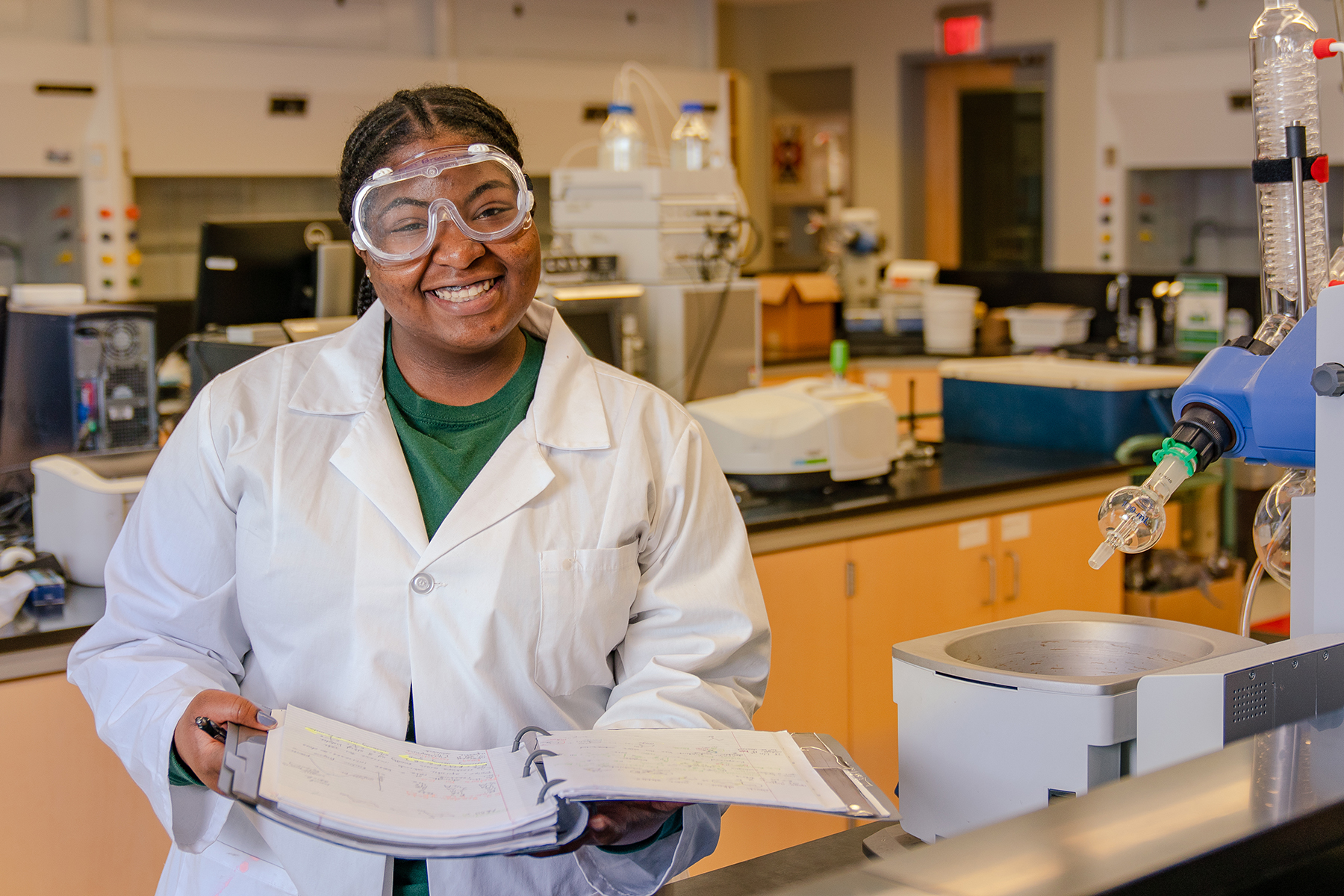

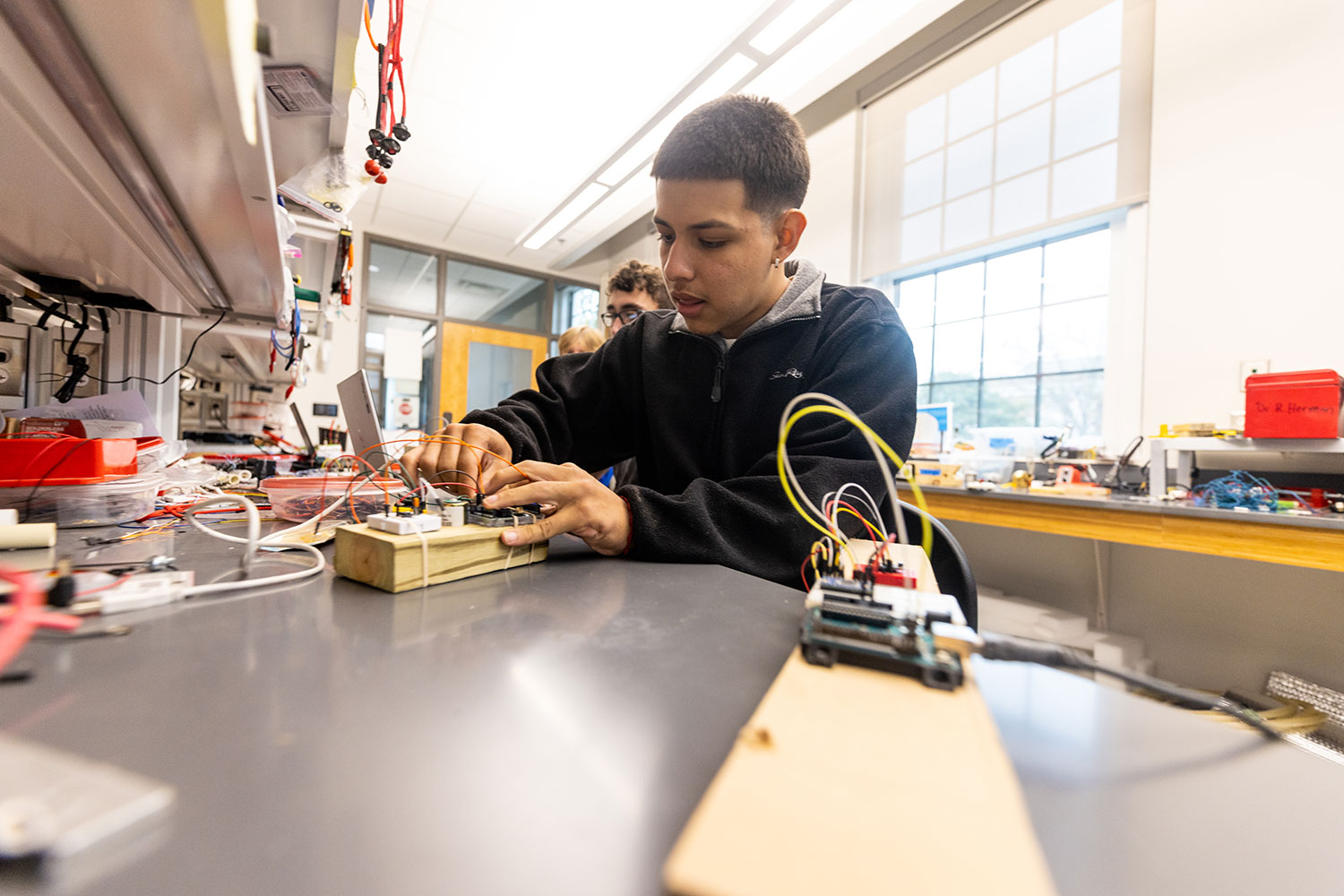
Breakthroughs in the science, technology, engineering and mathematics (STEM) fields are driving innovations that continuously transform our daily lives.
With this in mind, we prepare our students for successful careers in the STEM fields while actively engaging in real-world problem-solving.
Student Spotlight
Committees, Centers and Museums
The Artis College of Science and Technology at Radford University is home to a wide range of unique facilities that fosters curiosity, discovery, and practical experience across multiple scientific disciplines.
Academic Departments and School
News
-
Casting a kind eye on the ‘bad guy’
May 29, 2026
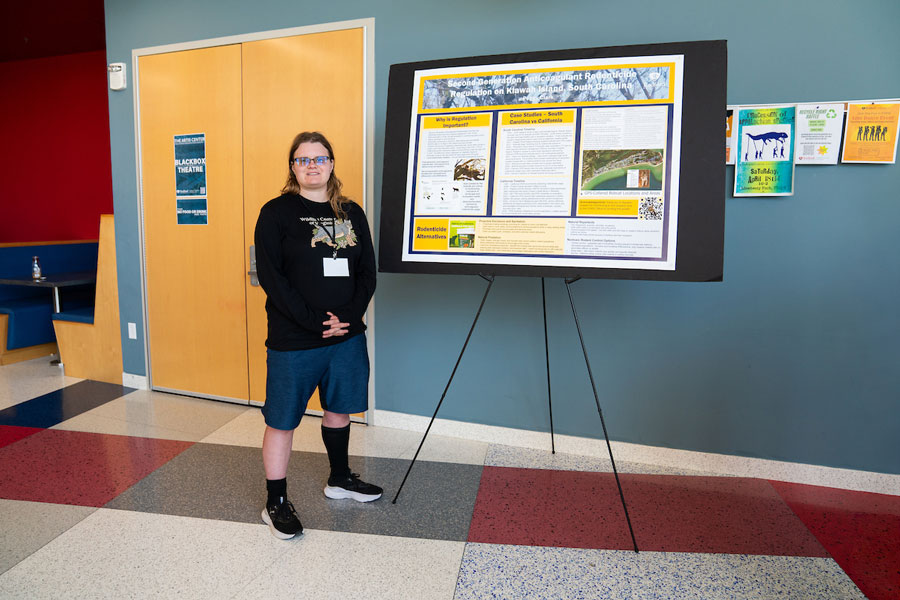
Senior biology major Peggy Clark has recently penned about a half dozen scholarly online articles that offer her conservation-minded perspectives on the typical ‘villains’ of the animal kingdom, the predators.

-
Professor, student explore research options for Highlanders in South America.
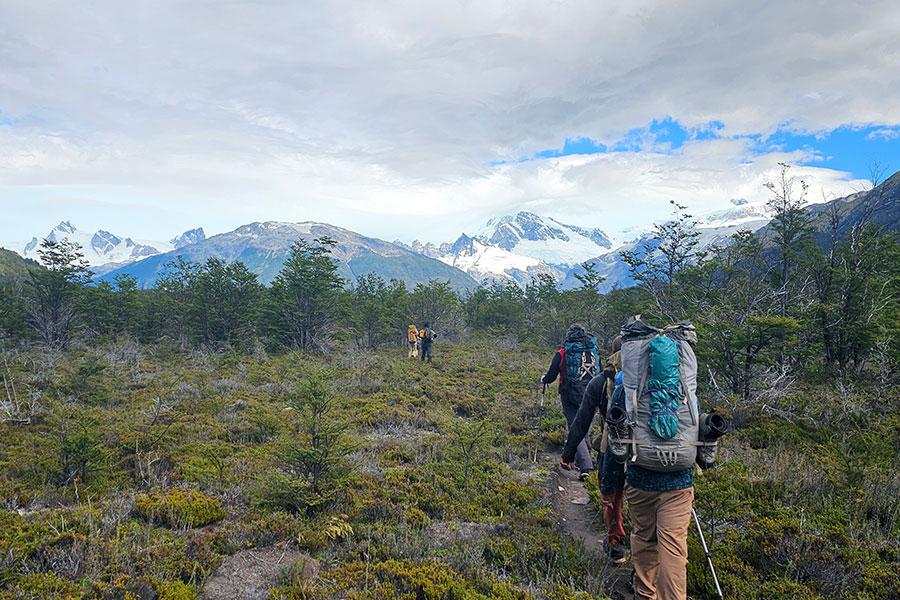

-
Radford University celebrated its Class of 2026 during Spring Commencement ceremonies held May 1-2, highlighted by an inspiring address from alumnus Eugene Naughton '89, president of The Dollywood Company.